

Liu Guoqing, Ph.D., Professor. He is a "Grassland Talent" of Inner Mongolia Autonomous Region and a second-level candidate of the "321 Talent Project" of Inner Mongolia Autonomous Region. He is also the executive director of the Inner Mongolia Society of Biotechnology and a director of the Inner Mongolia Society of Biophysics and Bioinformatics. Liu Guoqing was born in Kozhong Banner, Tongliao City, Inner Mongolia. He received his bachelor's degree in physics from Inner Mongolia Normal University in 2004 and his Ph.D. in biophysics from Inner Mongolia University in 2009. He visited the University of Georgia and Peking University for one year each in 2015 and 2019. Currently, he is employed at Inner Mongolia University of Science and Technology, where he is engaged in teaching and research in bioinformatics and biomedical big data mining. His research interests include tumor metabolic reprogramming, chromatin structure simulation, and the molecular mechanisms of meiotic recombination. He has led three projects funded by the National Natural Science Foundation of China and published over 50 academic papers. He has also won the First Prize of the Inner Mongolia Natural Science Award.
The occurrence and development of tumors are related to multiple factors. At the molecular level, in addition to gene mutations, chromatin epigenetic modifications and structural changes, abnormal tumor metabolism is an important driving force for tumor initiation and progression. Our research group focuses on intra-tumor heterogeneity and inter-tumor heterogeneity, the epigenetic regulatory mechanisms of metabolic reprogramming, the dynamic changes in metabolic pathways during tumor development, and the interaction mechanisms between metabolic reprogramming and the tumor microenvironment (such as the immune microenvironment).
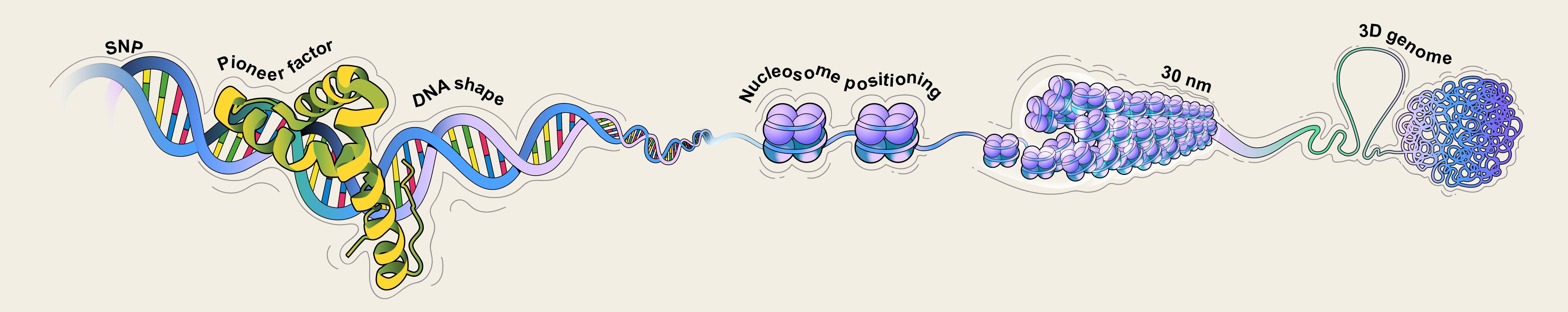
Eukaryotic chromatin has a multi-level structure: DNA double helices wrap around histone octamers to form nucleosome structural units, which are further stacked into a structure with a diameter of approximately 30 nm, and then folded into loop structures, forming the most compact structure during the metaphase of cell division. The hierarchical structure of chromatin regulates important biological processes such as gene transcription, chromosome replication, and chromosome recombination, thereby influencing cell differentiation and individual development. Our research group focuses on the dynamic changes in chromatin structure and their functions during cell differentiation, particularly through mathematical modeling of nucleosome structure and 30 nm structure to explore the regulation of chromatin structure by DNA sequence.
Meiotic recombination refers to the exchange of genetic material between homologous chromosomes during meiosis in eukaryotic cells. It is an important source of genetic diversity and the driving force for the adaptive evolution of sexual organisms. The chiasmata formed between homologous chromosomes during meiotic recombination are crucial for the correct separation of homologous chromosomes. Abnormal recombination is associated with genetic diseases, such as the occurrence of aneuploidy due to abnormal recombination. Meiotic recombination is a complex biological process involving the interaction of DNA sequence, chromatin structure, histone modifications, and multiple recombination-related enzymes. Our research group is interested in the following questions: (1) Studying the molecular mechanisms and regulatory networks of chromosome pairing and exchange from multiple dimensions, including sequence information of recombination hot and cold regions, nucleosome positioning, DNA methylation, histone modifications, and chromatin accessibility; (2) Investigating the evolutionary patterns of the genome under the influence of recombination; (3) Screening for genetic markers for parentage testing that are not affected by recombination; (4) Developing machine learning prediction models for recombination hot and cold regions.

Cang Jing, currently a master's student. She graduated from Inner Mongolia University of Science and Technology with a bachelor's degree in Bioinformatics and is now pursuing a master's degree in Biology at the same university. From January to March 2024, she interned at Hangzhou Zhijun High-tech Co., Ltd. Her current research interests include: screening genetic markers for paternity testing that are not affected by recombination, exploring tumor metabolic reprogramming, and building a team database.

Cong Rican graduated from Inner Mongolia University of Science and Technology with a bachelor's degree in Bioinformatics. During his undergraduate years, he was mainly responsible for building the database of the bioinformatics team. Currently, he works at Shanghai Lumine Biotechnology Co., Ltd. as a Metabolic Bioinformatics Engineer, mainly responsible for related metabolic analyses.