
Our lab focuses on cancer data-mining, chromatin structure modeling, and meiotic recombination, mainly via computational biology and bioinformatics. Our research is characterized by the thread of "omics data analysis — bioinformatics modeling — function elucidation". The main goal is to deepen the understanding of cancer development and assist cancer diagnosis, prognosis, and therapy by developing bioinformatics tools.
1. Metabolic reprogramming in cancer
In addition to molecular level factors like genome variation, epigenomic signals and signaling pathways, metabolic reprogramming plays an essential role in cancer development. We investigate inter/intra-tumor metabolic heterogeneity and dynamics of metabolic pathways during cancer progression, while developing diagnostic and therapeutic models.
2. Modeling chromatin structure
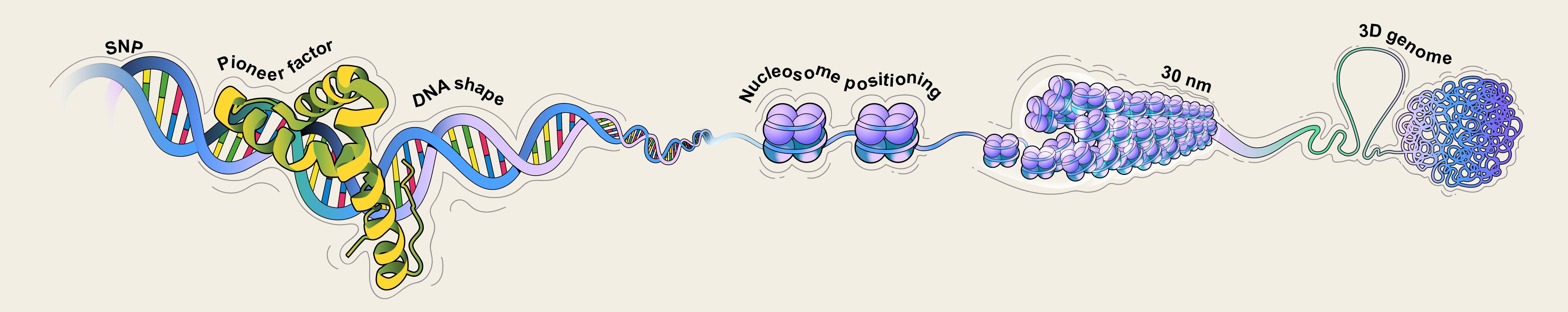
Studying multi-level chromatin structures (nucleosome, 30nm fiber, loops, TADs) and their regulatory roles in gene expression and cell differentiation. Focus on nucleosome positioning dynamics and cancer-related structural modeling.
3. Epigenetic regulation in meiotic recombination
Investigating recombination mechanisms regulated by epigenetic factors (DNA methylation, histone modifications, 3D genome). Developing machine learning models to identify recombination hotspots and study genome evolution.